|
It also provides the tools for basic analysis of protein sequences, DNA/RNA/protein alignment and DNA reads assembly. The program is designed to facilitate the cloning of restricted DNA fragments into the vector. BioLabDNA is the document based app focused on the analysis and manipulation of DNA sequences. This program is aimed at researches in the molecular biology field. The features of BioLabDonkey can be seen here – BioLabDonkey FeaturesīioLabDNA is one time purchase macOS program with lifetime updates including new features. It requires only the DNA sequences of the selected genes.Īcknowledgement I am grateful to Mike Dyall-Smith for help in developing the program and correcting the Help book. The method used for calculating RNA-seq differential expression between two samples can be performed for selected genes without alignment of reads to the genome or de novo transcriptome assembly. Users can also create their own custom databases from pdb files in addition to the default database, and this is most useful when the proteins of interest are similar to the proteins used to make the database, for example from pdb files of the same protein family, protein location (membrane/cytoplasm) or environment (halophilic, mesophilic, thermophilic). The default database of patterns was generated from publicly available pdb structures. The custom protein secondary structures prediction algorithm is based on patterns analysis of alpha helices, beta strands and turns. These custom algorithms are faster than the Needleman-Wunsch algorithm for large sequences having high homology. In contrast to the Needleman-Wunsch algorithm, the custom algorithms provide correct alignment independent of the size of the sequences indels. 
The program includes custom algorithms for: the protein/DNA sequences alignment, protein secondary structures prediction, RNA-seq differential expression. Analysis of Gel/Blot photo and calculation of relative fold-change in selected Gel/Blot bands/spots.Calculators for: Dilution, Molarity, Ligation, SDS PAG, Ammonium sulfate, Bacteria growth.RNA-Seq DE: genes and miRNAs differential expression.Find/highlight protease sites and protein fragments.Prediction of protein secondary structures.Multiple alignment and realignment of DNA/RNA/protein sequences.Find/highlight DNA restriction sites and DNA fragments.Copy/paste the selected DNA sequence with features.Generate DNA features database and perform features annotation.Generate and edit DNA/protein features for genbank/genpept format files.Generate Plasmid, Linear and Text map using feature annotations from genbank file.View GC content, CpG islands and Stem Loops.View ABI sequencing chromatogram, edit sequence. 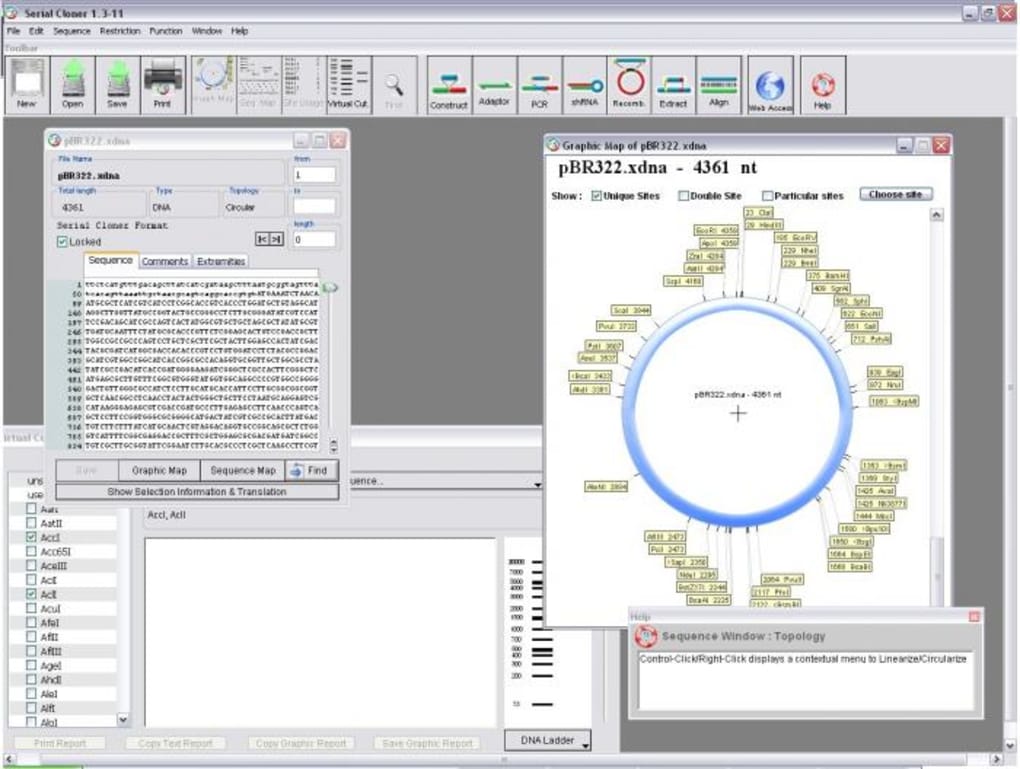
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
